A public beta of the GenePattern Notebook environment for JupyterLab is now available, and all are welcome to participate. To access the JupyterLab beta, go to https://lab.genepattern.org and sign in using your GenePattern credentials.
Feedback on the state of the beta is always welcome, and may be submitted using the public beta feedback form.
Getting Started
Within GenePattern Notebook for JupyterLab you will find a new "Toolbox" panel on the lefthand side of the interface. This panel may be used to browse, search and insert available bioinformatic tools, as well general machine learning methods. Simply open a new notebook, then click on the icon and select GenePattern Login to access the available bioinformatics analyses (see the figure below).
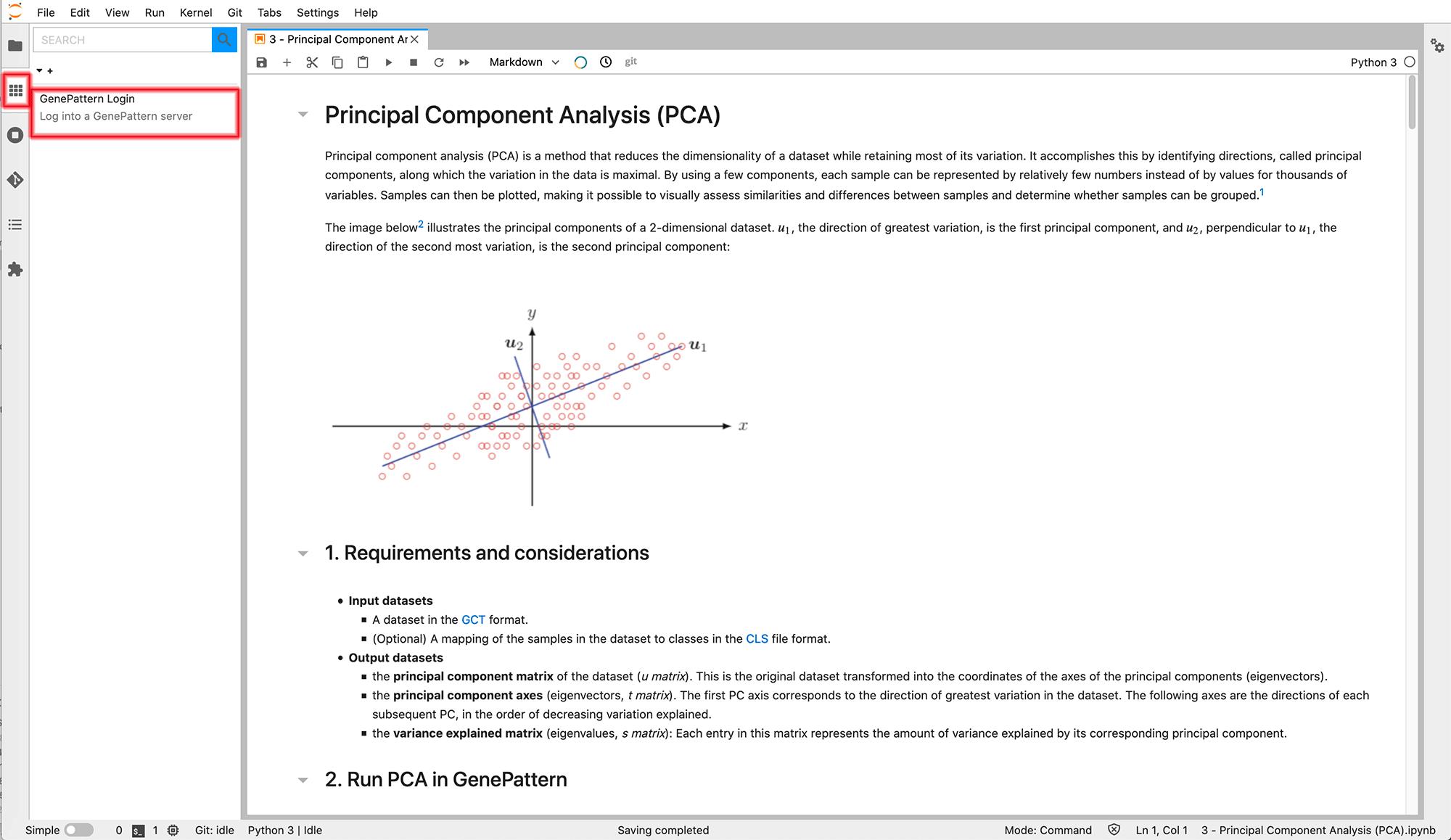
Once you insert a GenePattern Login cell, it will look like the widget outlined in the figure below. Simply enter your GenePattern credentials and click "Login to GenePattern". After you have logged in, you will be able to browse the full list of available analyses on the left.
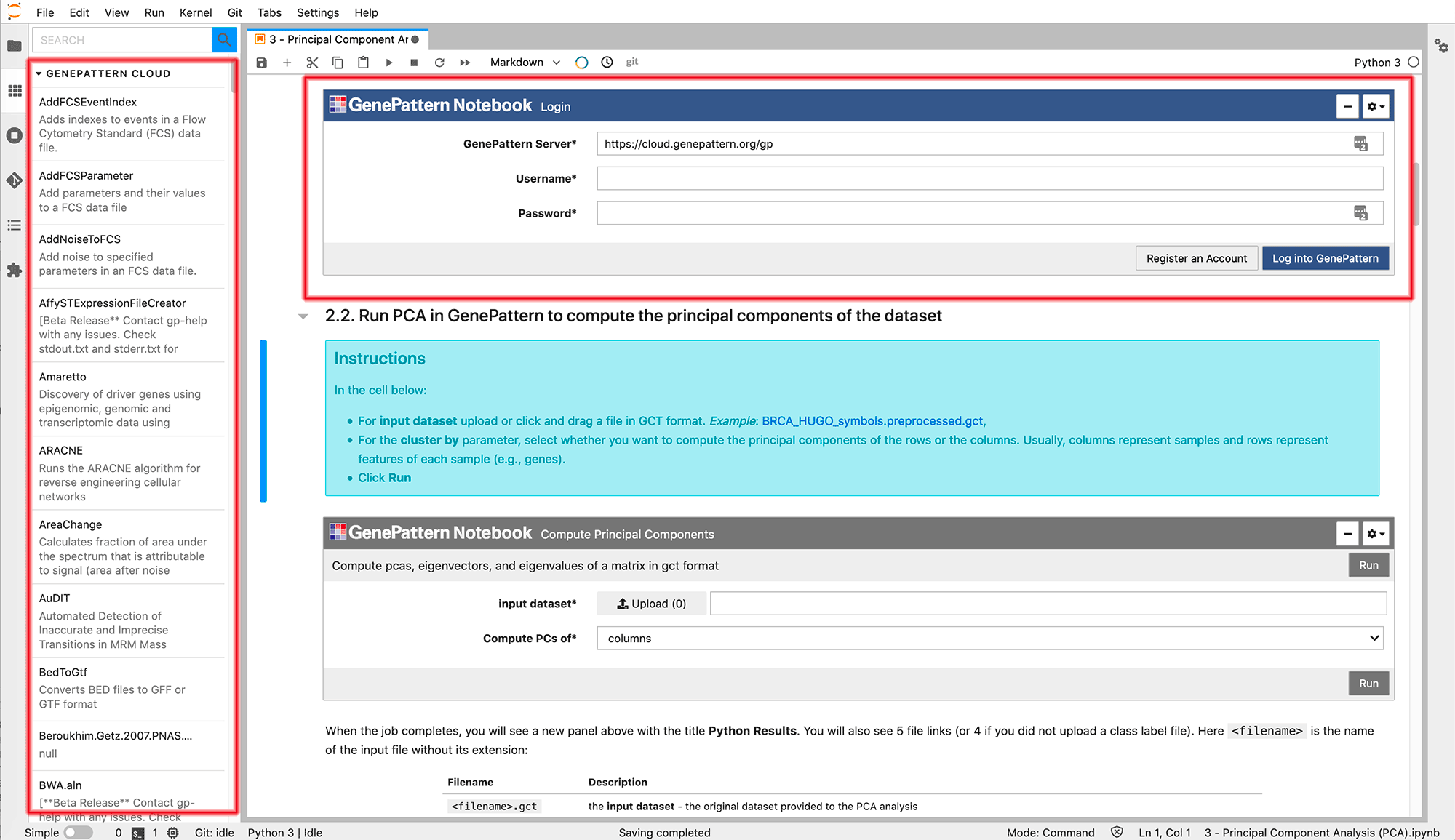
What is GenePattern Notebook?
The GenePattern Notebook environment provides an easy way to create and run a wide range of genomic analyses within shareable, electronic notebooks. Many common workflows are available for it in our Public Notebook Library. Included is the ability to seamlessly integrate GenePattern analyses with R and Python, but no programming or installation is required.
What is JupyterLab?
JupyterLab is a next-generation web-based user interface for Project Jupyter. It enables you to work with documents and activities such as Jupyter notebooks, text editors, terminals and custom components in a flexible, integrated and extensible manner. It includes synchronized visualizations and customizable panels, all in an easy-to-navigate tabbed interface.